In this tutorial we will download C3S Satellite Soil Moisture data from the Copernicus Climate Data Store (CDS), read data stacks in python using xarray and perform simple analyses of soil moisture anomalies over selected study areas.
The tutorial consists of the following main steps:
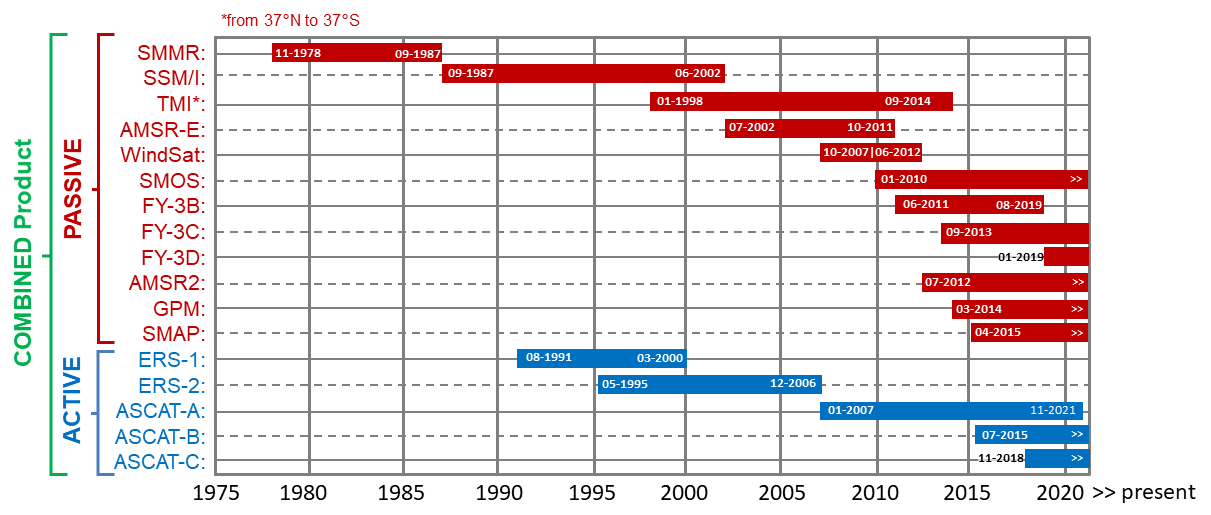
- A short description of the data sets
- Download satellite soil moisture images from CDS using the CDS API
- Application 1: Open downloaded data and visualize the soil moisture variable
- Application 2: Extract a time series for a single location and compute the soil moisture anomaly
- Application 3: Vectorized anomaly computation and data extraction for a study area
| Run the tutorial via free cloud platforms: |
|
|---|
Note: Cells in this notebook should be executed in order (from top to bottom). Some of the later examples depend on previous ones!